This function visualizes variable importance in models
Usage
varimp_plot(
model_list,
...,
type = "lollipop",
text_size = 10,
palette = "viridis",
n_row,
n_col,
save = FALSE,
file_path = NULL,
file_name = "VarImp_plot",
file_type = "pdf",
dpi = 80,
plot_width = 7,
plot_height = 7
)Arguments
- model_list
A
model_listobject from performingtrain_models.- ...
Additional arguments to be passed on to
varImp.- type
Type of plot to generate. Choices are "bar" or "lollipop." Default is
"lollipop."- text_size
Text size for plot labels, axis labels etc. Default is
10.- palette
Viridis color palette option for plots. Default is
"viridis". Seeviridisfor available options.- n_row
Number of rows to print the plots.
- n_col
Number of columns to print the plots.
- save
Logical. If
TRUEsaves a copy of the plot in the directory provided infile_path.- file_path
A string containing the directory path to save the file.
- file_name
File name to save the plot. Default is
"VarImp_plot."- file_type
File type to save the plot. Default is
"pdf".- dpi
Plot resolution. Default is
80.- plot_width
Width of the plot. Default is
7.- plot_height
Height of the plot. Default is
7.
Details
Note: Variables are ordered by variable importance in descending order, and by default, importance values are scaled to 0 and 100. This can be changed by specifying
scale = FALSE. SeevarImpfor more information.
Examples
# \donttest{
## Create a model_df object
covid_model_df <- pre_process(covid_fit_df, covid_norm_df)
#> Total number of differentially expressed proteins (8) is less than n_top.
#> None of the proteins show high pair-wise correlation.
#>
#> No highly correlated proteins to be removed.
## Split the data frame into training and test data sets
covid_split_df <- split_data(covid_model_df)
## Fit models based on the default list of machine learning (ML) algorithms
covid_model_list <- train_models(covid_split_df)
#>
#> Running svmRadial...
#>
#> Running rf...
#>
#> Running glm...
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#> Warning: glm.fit: algorithm did not converge
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
#>
#> Running xgbLinear...
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep1: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep1: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep1: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep1: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep1: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep1: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep1: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep1: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep1: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep1: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep2: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep2: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep2: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep2: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep2: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep2: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep2: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep2: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep2: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep2: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold01.Rep3: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold02.Rep3: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold03.Rep3: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold04.Rep3: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold05.Rep3: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold06.Rep3: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold07.Rep3: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold08.Rep3: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold09.Rep3: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=0e+00, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-01, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-04, alpha=0e+00, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=0e+00, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-01, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-04, alpha=1e-01, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=0e+00, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-01, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-04, alpha=1e-04, nrounds= 50, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=0e+00, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-01, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-04, alpha=0e+00, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=0e+00, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-01, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-04, alpha=1e-01, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=0e+00, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-01, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-04, alpha=1e-04, nrounds=100, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=0e+00, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-01, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-04, alpha=0e+00, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=0e+00, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-01, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-04, alpha=1e-01, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=0e+00, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-01, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: Argument 'objective' is only for custom objectives. For built-in objectives, pass the objective under 'params'. This warning will become an error in a future version.
#> Warning: model fit failed for Fold10.Rep3: lambda=1e-04, alpha=1e-04, nrounds=150, eta=0.3 Error in modelFit$xNames <- colnames(x) :
#> ALTLIST classes must provide a Set_elt method [class: XGBAltrepPointerClass, pkg: xgboost]
#> Warning: There were missing values in resampled performance measures.
#> Something is wrong; all the Accuracy metric values are missing:
#> Accuracy Kappa
#> Min. : NA Min. : NA
#> 1st Qu.: NA 1st Qu.: NA
#> Median : NA Median : NA
#> Mean :NaN Mean :NaN
#> 3rd Qu.: NA 3rd Qu.: NA
#> Max. : NA Max. : NA
#> NAs :27 NAs :27
#> xgbLinear failed.
#>
#> Running naive_bayes...
#> Done!
## Variable importance - lollipop plots
varimp_plot(covid_model_list)
#> Warning: The `size` argument of `element_line()` is deprecated as of ggplot2 3.4.0.
#> ℹ Please use the `linewidth` argument instead.
#> ℹ The deprecated feature was likely used in the promor package.
#> Please report the issue at <https://github.com/caranathunge/promor/issues>.
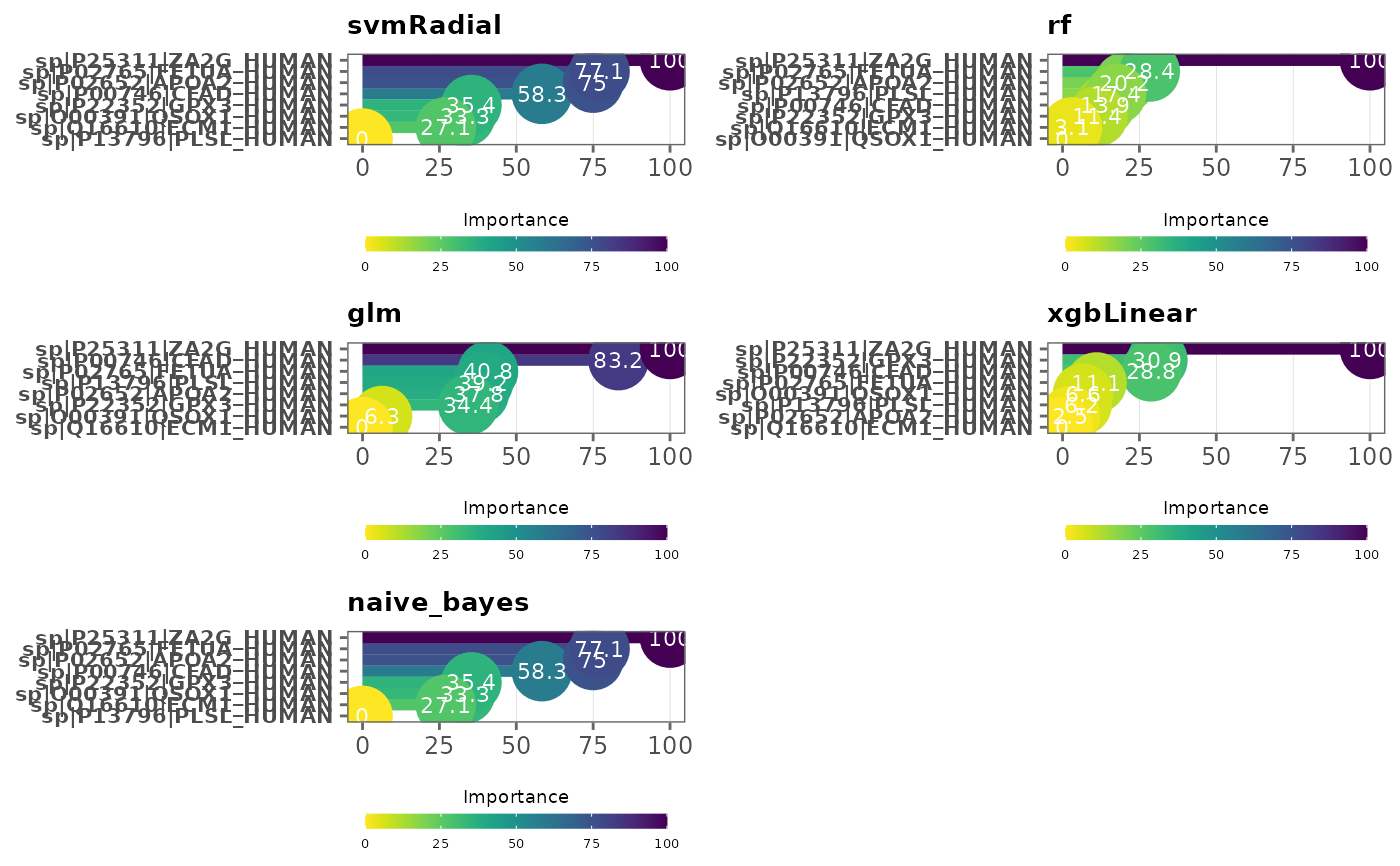 ## Bar plots
varimp_plot(covid_model_list, type = "bar")
## Bar plots
varimp_plot(covid_model_list, type = "bar")
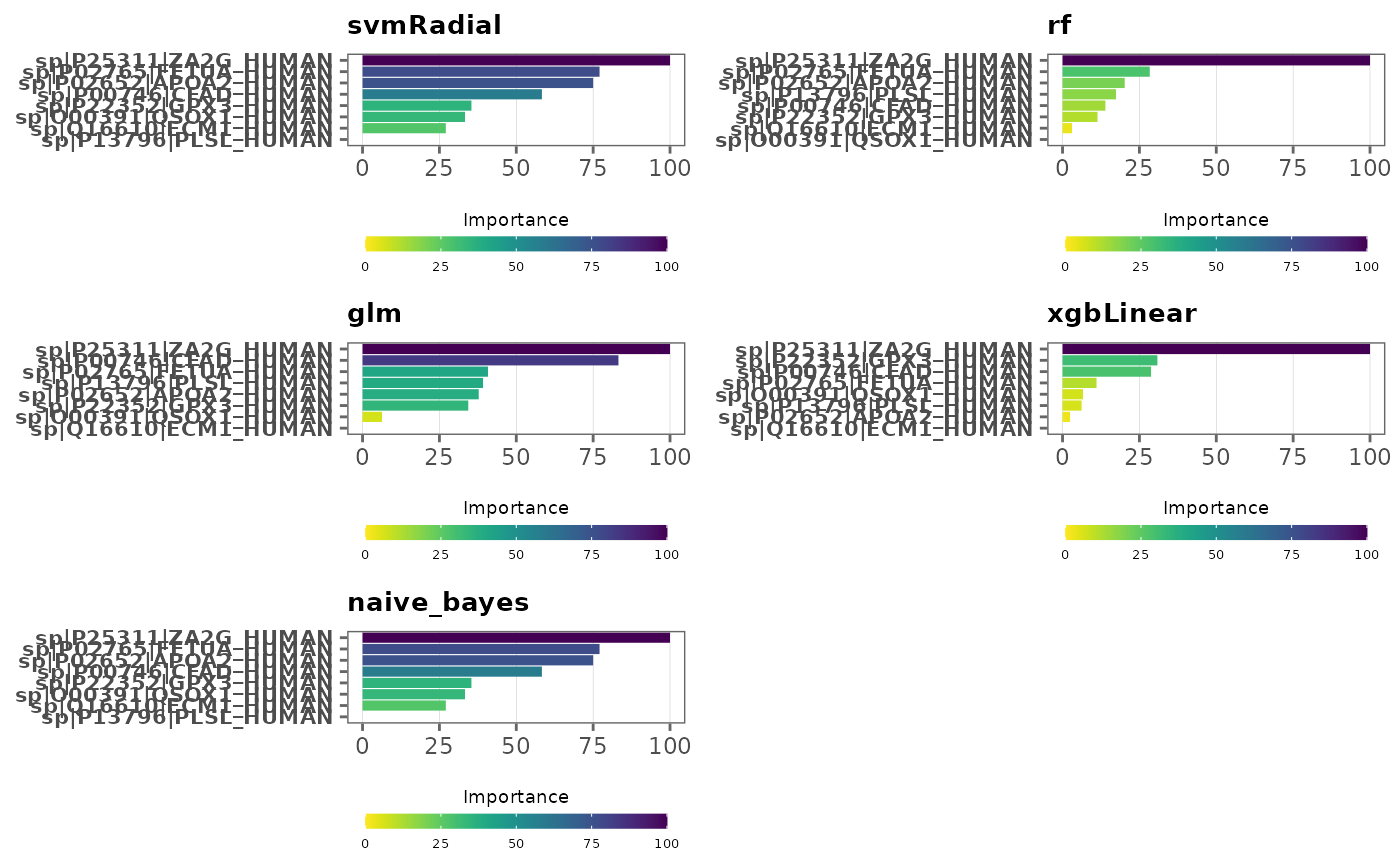 ## Do not scale variable importance values
varimp_plot(covid_model_list, scale = FALSE)
## Do not scale variable importance values
varimp_plot(covid_model_list, scale = FALSE)
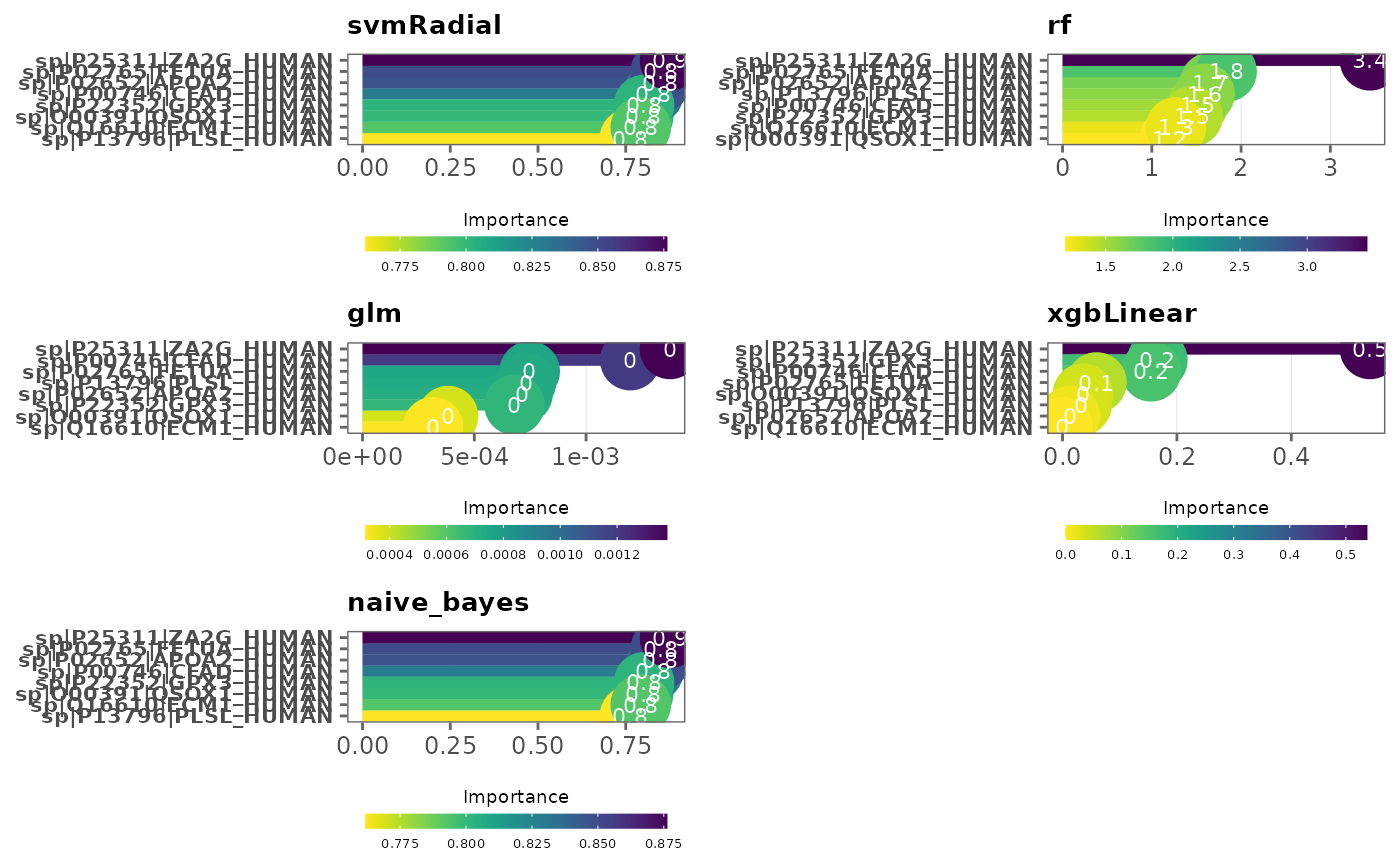 ## Change color palette
varimp_plot(covid_model_list, palette = "magma")
## Change color palette
varimp_plot(covid_model_list, palette = "magma")
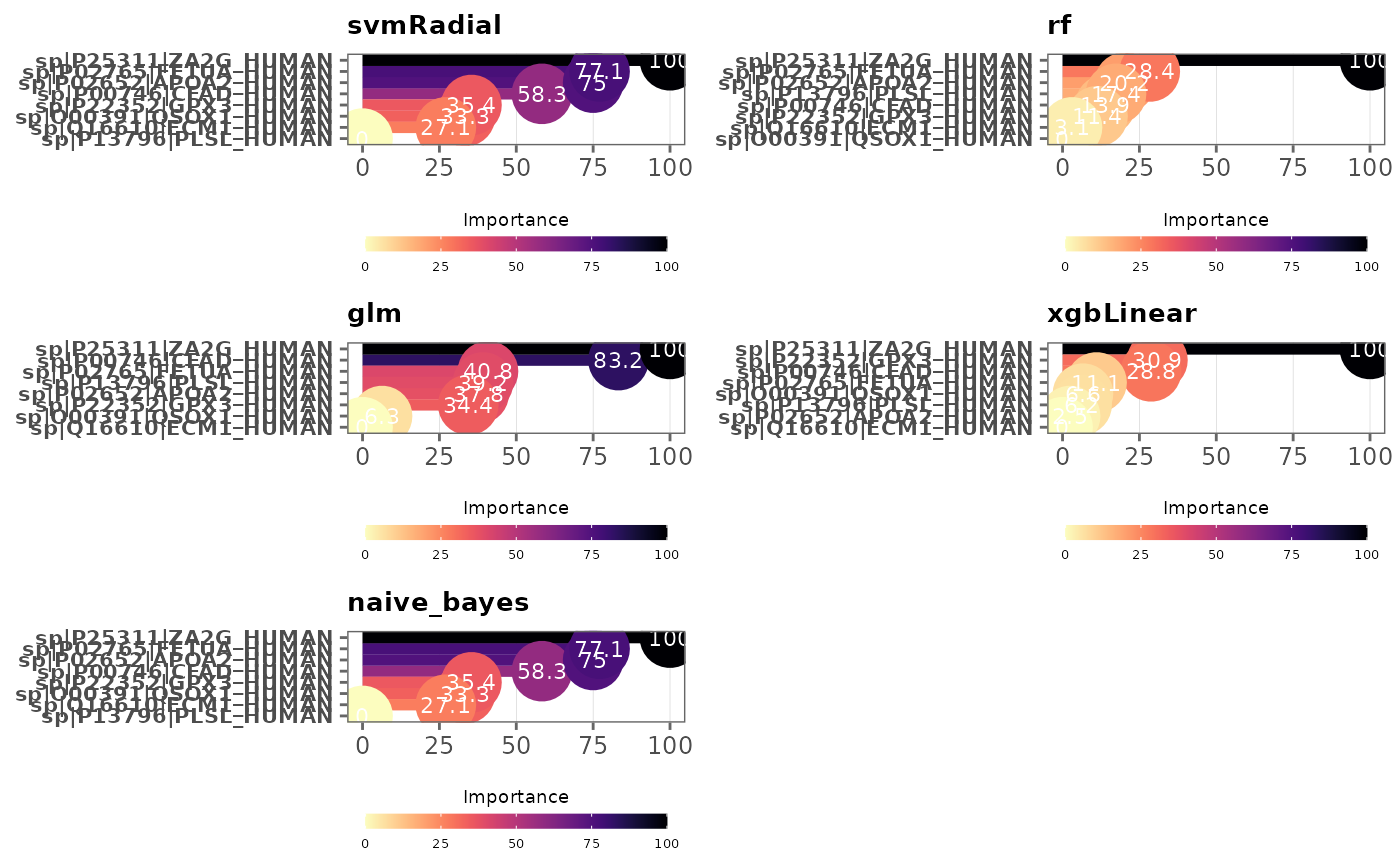 # }
# }
